Protein Three-Dimensional Visualization Assignment
CHEM-405 Biochemistry I
Adapted from "How to visualize, understand and share/present 3D protein molecular structures"
by Eric Martz, 2016
I. Getting Started with the Software
Firefox, Chrome, Safari, Edge work well. Opera works in Windows but not Mac. DO NOT USE Internet Explorer.
- Go to Atlas.MolviZ.Org.
- In the Atlas, select a category of molecules (inside the yellow part of the page) and then select any protein listed as STRAIGHTFORWARD (not one in the CHALLENGING list!) in that category and click on the link to FirstGlance. After a few seconds to load (depending on your connection speed), you should see a rotating molecule.
- If you have any difficulty or the molecule does not appear, ask for help!
- Note that this enzyme will be one you'll just use to familiarize yourself with the software. It won't necessarily be the one you'll use for your enzyme paper, as you'll select that enzyme in part IV.
II. Goals
1. Review principles of protein 3D structure.
2. Choose an existing experimentally-determined 3D protein structure model to investigate for your Enzyme Paper Project.
3. Learn how FirstGlance in Jmol makes it easy to see structure-function relationships in the protein you chose.
4. Write a PowerPoint slides report including snapshots of your protein (future images for your paper!) that illustrate your answers to the questions below, which you'll upload to Canvas.
III. Protein Structure and Structural Bioinformatics
|
|
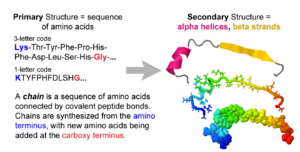
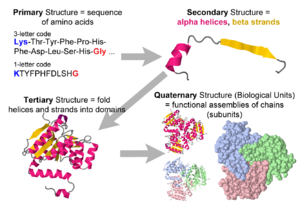
- 1. Amino acid sequence + protein chain conformation = protein function.
- A. Why do we care about protein 3D structure?
- B. Conformation can be a stable fold or intrinsically unstructured. Both commonly exist in the same protein molecule.
- C. Conformation is specified by sequence.
- Folded domains fold spontaneously (Anfinson, 1960's[1]), or with the help of chaperonins.
- The denaturation (unfolding) of a folded protein domain destroys its function.
- 2. Backbone Representation.
- A. Backbone Representations
- B. Small Protein in FirstGlance (use the Views Tab: Vines, Cartoon)
- 3. Structure Knowledge.
- A. Although sequence specifies fold, scientists only recently became able to predict the fold from the sequence (See AlphaFold!). Therefore, fold must be determined by empirical (experimental) methods. The most common methods for determining the 3D structure of a protein molecule are:
- X-ray crystallography, 88%.
- Result is a single model representing the average of the molecules in the crystal.
- Resolution reflects the degree of order or disorder in the crystal.
- X-ray crystallography gives no models for intrinsically unstructured loops or molecules.
- Nuclear magnetic resonance (NMR) in aqueous solution, 9%.
- NMR is limited to small proteins (30 kD or smaller; median NMR in PDB is 10K; median X-ray is 50K).
- Result is an ensemble of models consistent with the data. Examples: 2bbn
- A. Although sequence specifies fold, scientists only recently became able to predict the fold from the sequence (See AlphaFold!). Therefore, fold must be determined by empirical (experimental) methods. The most common methods for determining the 3D structure of a protein molecule are:
- B. These methods are difficult and expensive. Less than 10% of proteins have known structure (until AlphaFold, that is!).
- C. All published, empirically determined 3D macromolecular structure models are available from the Protein Data Bank (PDB; pdb.org; About the PDB).
- E. Crystallographers publish the asymmetric unit of the crystal. It may be identical with the biological unit (the functional form of the molecule), or it may be only part of the biological unit, or it may contain multiple copies of the biological unit. See examples.
- Interchain contacts that occur in the asymmetric unit, which are absent in the biological unit, are an artifact of crystallization, termed crystal contacts.
- E. Crystallographers publish the asymmetric unit of the crystal. It may be identical with the biological unit (the functional form of the molecule), or it may be only part of the biological unit, or it may contain multiple copies of the biological unit. See examples.
IV. Choose an Enzyme to Explore
- Look for PDB codes and use FirstGlance to view them. Here are some of the ways to find PDF codes for proteins with known structure:
- RCSB Protein Data Bank, searching by molecule name or amino acid sequence.
- Atlas.MolviZ.Org. Choose a "Straightforward" or "Challenging" (not "Enormous") molecule that has protein and ligand (co-factor).
- Topic Pages in Proteopedia, or its Table of Contents.
- Random PDB Entry in Proteopedia.
- Choose an enzyme to use for your report. (Note that this enzyme will likely also be the one you use for the Enzyme Paper Project, so it's probably worthwhile finding one that interests you!)
- It must be a protein.
- The structure must be experimentally-determined (ie. X-ray crystal, NMR, or Cryo-EM) and not a computed structure model (ie. predicted by a computer).
- It must catalyze a chemical reaction with one or more reactants and products.
- It does not have to be listed at Atlas.MolviZ.Org
- You must each choose a different enzyme.
- Make sure it has at least one cofactor, as it will be more interesting if it contains some bound, non-protein cofactor, which can be either organic (eg. flavin), inorganic, (eg. Zn2+), or both (eg. heme).
- X-ray results should have resolution of 3 Å or better (less).
- Be sure to note the 4-character PDB code of the enzyme you choose. The PDB code makes it easy to retrieve the enzyme and information about it.
- Check the PDB code you choose with others posted to the Discussion page on the Canvas website to make sure it is not already taken.
- Post the PDB code, the enzyme name, and your name to the Discussion thread page on Canvas. First to choose a protein gets to use it!
V. Explore Your Enzyme
FirstGlance in Jmol
The main tool we will use is FirstGlance in Jmol[2]): FirstGlance.Jmol.Org. (To google it later, use the single word (no space) firstglance.)
- Enter your 4-character PDB code at FirstGlance, and you should see the enzyme you have chosen.
- Get familiar with what the molecule information tab tells you. Ask about anything you don't understand.
- In the Views tab, there are 10 links at the top that show you different aspects of the molecule. Try them all, as well as any of the other tools in FirstGlance that interest you.
-
FirstGlance does NOT use Java unless you tell it to. Using Java will make larger proteins load faster, rotate more smoothly, and change views quicker. Chrome does not support Java. In your Java-compatible browser, display a molecule in FirstGlance, and then click on the Preferences tab in FirstGlance. See Installing and enabling Java.
Here are two Views in FirstGlance that will be used in your report:
A. Hydrophobic/Polar
- Water-soluble proteins have polar/charged amino acids nearly everywhere on their surfaces (Examples: small 2hhd, large 1igy). Patches of hydrophobic amino acids on the surfaces of soluble proteins are usually less than ~10 å in their smaller diameter, and usually recessed.
- Hydrophobic surface patches may be buried in chain-to-chain contacts -- check the biological unit (example: lac repressor homodimer).
- Large, protruding hydrophobic surface areas (>25 Å in their smaller diameter) may indicate transmembrane proteins (insoluble). Examples:
- 7ahl
- showing bilayer boundaries (click on "Jmol"; ligand toggles boundaries).
- 1bl8
- Gramicidin Channel in Lipid Bilayer.
B. Charge
Most proteins have roughly equal numbers of positive and negative charges intermixed on their surfaces. Surface patches of exclusively positive charge often bind nucleic acids (negatively charged because of their phosphates). For example, examine the protein surface charges where the gal4 transcriptional regulator binds DNA (1d66).
VI. Report Slides
Answer the questions below in PowerPoint slides.
- Name your report YourLastName_YourFirstName_PDBCode, for example Osborne_Jeff_4D7B. If the name of your report does not begin with your family name, you will lose 2 points. When completed, upload your report to Canvas for grading.
- You will not be asked to present this report in class.
- Each slide MUST be labeled at the top with its section number, e.g. Section 1.
- Each Section below may be answered in a single slide, or multiple slides. For example, suppose you want to show two snapshots for Section 3, and make separate comments. You may choose to use two slides, labeled Section 3A and Section 3B.
- Additional work beyond the minimum required may earn extra credit.
- Due date is listed in the syllabus.
This is not an exam. It is to help you learn by doing. Ask for help!
Example of a Completed Report (You may import these slides into a new presentation of your own, and use them as a templates, putting in your own content.)
Section 1: Identity
- The label Section 1 at the top (and so forth for every slide).
- Your name.
- Your PDB identification code.
- The name of your enzyme.
- The function of your enzyme (briefly, one sentence).
- The experimental method and resolution (or number of models for NMR). Available in the molecule information tab in FirstGlance.
- A snapshot of your enzyme.
is also linked at the bottom left in FirstGlance. |
Section 2: Composition
- The number of
- Protein chains (What does "chain" mean?)
- DNA chains
- RNA chains
- Available in the molecule information tab in FirstGlance: Chain Details. You can identify a residue in any chain by touching it with the mouse (spinning off!). DNA residues are DA, DG, DC, DT while RNA residues are A, G, C and U.
- Ligands (Co-factors) and Non-Standard Residues: Give the one to three-letter abbreviations and full names for all ligands (co-factors) and non-standard residues. If none, so state. (Standard residues)
- The molecule information tab in FirstGlance lists the 1 to 3-letter abbreviations for each ligand (co-factor) and non-standard residue, and their full names. Click on an abbreviation to locate that entity in the model. See also Composition in FirstGlance's Views tab.
Section 3: Evolutionary Conservation
Go to How to see conserved regions, where it will direct you to enter your PDB code in Proteopedia to see conserved residues.
Does your enzyme have a highly conserved region? If so, what is its function? If there is no highly conserved region, is there a highly variable region? Show two snapshots illustrating a highly conserved region, and a contrasting region.
See How to see conserved regions.
- If Proteopedia lacks a pre-calculated Evolutionary Conservation for your enzyme, and you do your own calculation at the ConSurf Server, be sure to include the address of the ConSurf result in your report slide!.
Section 4: Hydrophobic/Polar
Do you think your enzyme is water soluble? Support your conclusion with a snapshot. Be sure to use the Hydrophobic/Polar view from FirstGlance in a snapshot. Optionally, you may show other views in other snapshots.
Section 5: Hydrophobic Core
Are there hydrophobic cores in your enzyme? For soluble proteins, expect a hydrophobic core in each domain. Support your conclusion with a snapshot. Be sure to use the Hydrophobic/Polar view from FirstGlance and turn on the Slab button.
Section 6: Charge
Are there any areas on the surface of your enzyme with only positive (or negative) charges? Show snapshots illustrating your conclusions. Be sure to use the Charge view from FirstGlance in your snapshots.
Section 7: Cation-Pi Interactions
Show a snapshot of an energetically significant cation-pi interaction. Include a distance monitor in your snapshot. Also paste in the report from CaPTURE confirming its energetic significance. The cation-pi interaction tool, and instructions for measuring distances, are in the Tools Tab. If you are having trouble getting the CaPTURE website to find your structure (mostly blank page appearing), download the .pdb file for your structure (available at https://www.rcsb.org/ and other places) and then upload that .pdb file onto the CaPTURE website.
Section 8: Biological Unit
In FirstGlance, in the molecule information tab click Biological Unit. (It is also in the Resources Tab.)
How many total polymer chains (protein + DNA + RNA) are in the asymmetric unit? The Biological unit?
Show side-by-side two snapshots comparing the asymmetric unit with the biological unit. The Cartoon representation in FirstGlance is best for these snapshots. Make sure to label which is which.
Section 9: Animation from Polyview-3D
Minimal steps to make an animation:
- Go to Polyview-3D.
- Enter your PDB code in the PDB ID slot near the top. (If the slot is not visible, open the section Source of Structural Data.)
- Change "Type of request" from "Single slide" to "Animation". It is under the Image Settings section near the bottom.
- Click any "Preview" link.
- Optional: If you want to modify the orientation or zoom of the molecule, click on View by Jmol / Set orientation under Quick links at the upper left of the page. Use the mouse to rotate and zoom in Jmol. Then click the Set and close button.
- Optional: If you want to change the colors, hide portions of the molecule, emphasize certain residues, etc. feel free to try out these options in the form, using Preview to check your results.
- In the "Animation Settings" section at the bottom of the page, set Delay to 10/100.
- Change "Angle step" to 5 degrees.
- Check "Rocking".
- Change "angle range" for rocking to 30 degrees.
- Click "Submit".
The above steps are the minimum for an animation that avoids putting a heavy load on the server. Feel free to try other options, but while everyone else in the class is simultaneously making an animation, please don't make a large (>300 pixel) animation, or increase the angle range, or decrease the angle step size. Otherwise, the server may get overloaded and take a very long time to produce results. When others aren't using the website server, feel free to submit more demanding jobs. If you highlight specific residues, please explain why.
After your animation is completed and appears in the PolyView-3D web page:
- Right-click (Mac: control-click, or trackpad 2-finger click) on the animation in the Polyview-3D web page, and select Save Image As ...
- Save the image to the Desktop.
- Drag the image file (filename ending in .gif) from the Desktop and drop it into a slide.
- In PowerPoint SlideShow view, the animation should move immediately.
Examples of Slides with Polyview-3D Animations (These slides are only to show you what is possible. These are not in your assignment.)
Section 10 - Contacts/Non-covalent Bonds
Example: 4d7b.
- Click Contacts & Noncovalent Interactions in the Tools Tab in FirstGlance.
- Change target selection to Residues/Groups.
- Here one can click on something small to select it as a "target", such as a ligand (co-factor), or a single amino acid. Choose an amino acid with an uncharged polar side chain, such as Ser, Thr, Asn, Gln, Tyr, His.
- Click the link to Show atoms contacting target.
- Click Center contacts.
- Uncheck Backbones.
- Click the 4th thumbnail image
 to display the contacts as balls and sticks colored by element. The element color key is at the bottom of the Contacts help panel in FirstGlance.
to display the contacts as balls and sticks colored by element. The element color key is at the bottom of the Contacts help panel in FirstGlance.
- Zoom in (and click Return to Contacts if necessary).
- Uncheck all categories of non-covalent bonds.
- Check hydrogen-bonded non-water. (Review hydrogen bonds.)
- Double click the hydrogen bond donor and acceptor atoms to insert a distance monitor.
Describe the moiety (component) you selected as a target. Include a snapshot showing exactly one hydrogen bond. Be sure to identify the two entities (amino acids, nucleotides, ligand) by name, chain, and sequence number. I need enough detail to be able to reproduce what you are reporting.
Section 11 - How Structure Supports Function
Write a brief description in your own words (avoid plagiarism!) of how the structure of this protein supports its function. Doing some online research will strengthen your description.
Include links to supporting references. Use authoritative sources, such as peer-reviewed scientific journal articles or government websites (of course not Wikipedia).
Your description should be at least 75 words. More work, if well done, will earn more credit.
Section 11 in the Examples of a Completed Report is longer than the minimum required, but illustrates the sort of thing that could be done if you can spend the time.
VII. Notes and References
- For a brief overview of Anfinson's protein folding experiments in the 1960's, see the first paragraph at Intrinsically Disordered Protein.
- Cite FirstGlance in Jmol with the following two references: "Version 2.73 of FirstGlance in Jmol by Eric Martz is licensed under a Creative Commons Attribution-Noncommercial-Share Alike 3.0 United States License." and "Martz, E. 2005. FirstGlance in Jmol. http://firstglance.jmol.org"